Transmission tree reconstruction with SCOTTI
What you will need:
-BEAST 2.5.0 or greater
-Tracer 1.7.0 or greater
-FigTree
-Python 2.7 or greater
Transmission tree reconstruction with SCOTTI
In this tutorial we will use SCOTTI (Structured COalescent Transmission Tree Inference) to reconstruct transmission trees for small outbreaks (De Maio et al., 2016). A great tutorial introducing SCOTTI has already been created by Louis du Plessis and Nicola de Maio, so we will follow along with their tutorial available on the Taming the BEAST website:
I recommend downloading/cloning the entire repository for the tutorial so you will have all the data files and helper scripts in the same place.
IMPORTANT NOTE: Unfortunately, newer versions of BEAST 2 (>2.7) and the XML files created by the python script in the SCOTTI tutorial are incompatible. However, there are two easy workarounds:
One solution is to run an older version of BEAST 2. Any of the 2.5.x versions should be compatible and are available on the BEAST 2 releases site. However, to run these older versions of BEAST 2 you may need to also install an older version of the Java RunTime Environment.
The other solution (recommended) is to download a newer example of the SCOTTI XML input file. You won’t be missing out on much if you choose this option, as the original tutorial uses a Python script to generate these XML files for you. Here are links to newer XML files for the FMDV and Klebsiella datasets:
Reconstructing a neonatal Klebsiella pneumoniae outbreak
In addition to the FMDV dataset the Taming that the BEAST tutorial explores, De Maio et al. (2016) analyzed an outbreak of antimicrobial resistant K. pneumoniae in a Nepali neonatal intensive care unit. This was a severe outbreak that resulted in the deaths of 16 out of 25 infected infants. As an alternative to the FMDV outbreak dataset, you can follow along with the Taming the BEAST tutorial while using the files below to replicate the K. pneumoniae analysis. Note, however, that this analysis will take quite a bit longer to run since this it is a larger outbreak with more sampled hosts.
If you cloned the SCOTTI-Tutorial repo, you can place these files in the ‘Data’ folder alongside the FMDV data. You should then be able to run the SCOTTI_generate_xml.py as described in the tutorial, but replace the file name arguments with the K. pneumoniae file names as in the command below:
python SCOTTI_generate_xml.py --fasta ../data/KPneu.fasta --dates ../data/KPneu_dates.csv --hosts ../data/KPneu_hosts.csv --hostTimes ../data/KPneu_hostTimes.csv --output KPneu --maxHosts 40 --numIter 4000000 --tracelog 2000 --treelog 20000 --screenlog 20000
Running the python script should generate a KPneu.xml file that you can run in BEAST. Note that we’ve also increased the –maxHosts argument to 40 since we know that at least 25 hosts were infected. You should then be able to follow along with the rest of the Taming the BEAST tutorial without any problems.
After processing the BEAST output, I get a MCC tree that looks like this:
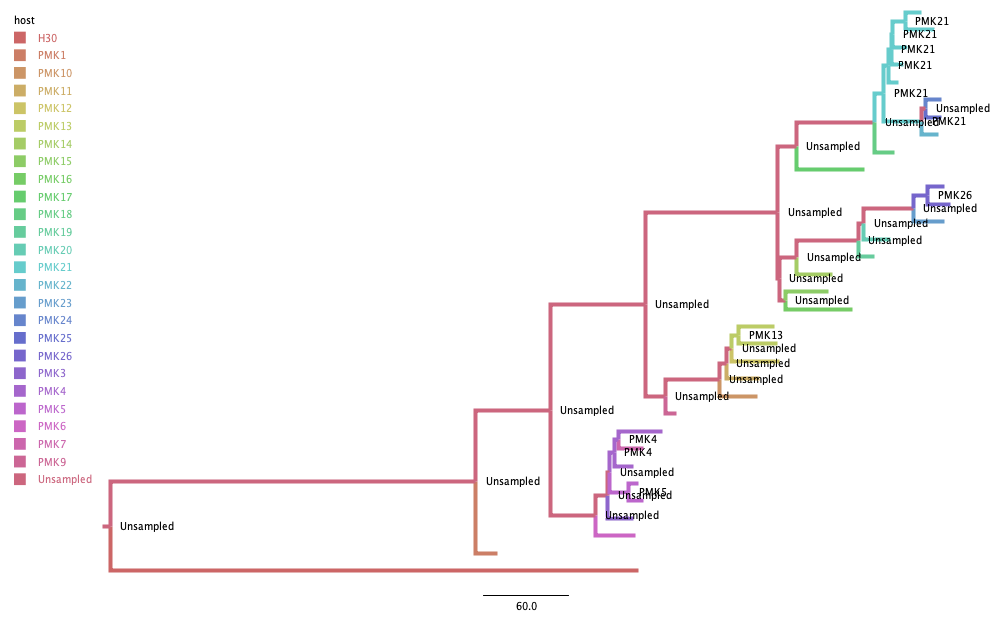
As in De Maio et al. (2016), most infections appear to be linked by unsampled hosts.